In Leo Breiman's legendary paper on
"Statistics modeling: the two cultures" a trend in statistical research was criticized: people, holding on to their models (methods), look for data to apply their models (methods). "
If all you have is a hammer, everything looks like a nail," Breiman quoted.
A similar research scheme was observed recently in data science. The availability of large unstructured data sets (i.e.,
Big Data) have sparkled imagination of quantitative researchers (and data scientists wanna-be) everywhere. Challenges such as the
Linked-In economic graph challenge have invited people to think hard of creative ways to unlock the information hidden in the vast "data" that can potentially lead to novel
data products. The exploratory nature of this trend, to some extent, resembles the quest of a hammer in search for nails. Only this time, it is a vast facet of under-utilized wall in search of deserved nails. Once the most deserved nails (or hooks and other wall installations) are identified, the most appropriate hammers (or tools) will be identified or crafted for the installation of them.
In every data science project, there is this trinity of three basic components: the
problem to be addressed, the
data to be used and the
method/model for the hack.
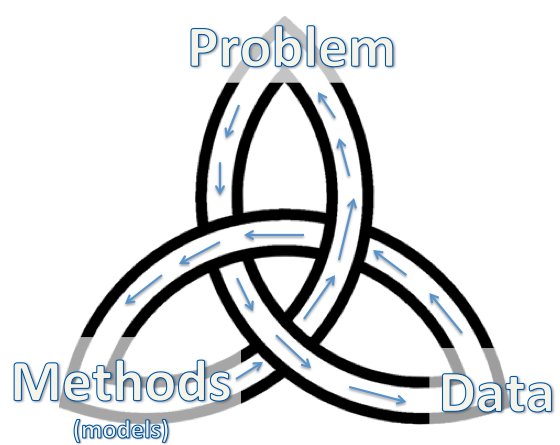 |
| The Trinity of Data Science |
Traditional statistical modeling usually starts with the problem, assuming certain generative mechanism (model) for a potential data source, and device a suitable method. The hack sequence is then
problem-data-methods (or first start with the nail, then choose which wall to use, and then decide which hammer to use, considering the nature of the nail and the wall).
Data mining explores data using suitable methods to reveal interesting patterns and eventually suggests certain discoveries that addresses important scientific problems. The hack sequence is then
data-methods-problem (the wall, the available hammers, and the nails).
Data scientists enter this tri-fold path at different points, depending on their career path. The ones from an applied domain have most likely entered from a problem entry point, then to data and then to methods. It is often very tempting to use the same methods when one moves from one set of data to the next set of data. The training of these application domain data scientists often comes with a "manual" of popular methods for their data. Data evolve. So should the adopted methods, especially given the advancements in the methodological domain. The hammer used to be the best for the nail/wall might no long be the best given the current new collection of hammers. It is time to upgrade.
Methodological data scientists such as statisticians enter from the methodological perspective due to their training. When looking for ways to apply or extend their methods, they should consider problems where their methods might be applied and then find good data for the problem. In the process, one should never take for granted that the method can just be applied to the problem-data duo in its original form.
Computational data scientists and engineers often started from manipulating large data sets. These Algorithms were motivated by previous problems of interests or models that have been studied. When a similar large set of data become available, the most interesting problems can be answered by this set of data might be different from the ones have been addressed before in another data set. One should use creative methods to identify novel patterns in such data and discovery interesting problems to answer.
Looking at this trinity map of data science, it is then easy to understand that, for some, there will naturally be phases when one knows a few methods (from their training, or prior hacks of data sets) and looks for (other) data sets or problems to hack; and for some others, there will be phases when one has a big data set and looks for problems that can be answered by this data set. And there will also be those who start with a problem, find or collect data and apply existing or novel methods on the data.
These are all valid and natural "entry points" into data science. The most important thing here is that one remembers that there are many different hammers, many different nails and many different walls. A quest of a data scientist shall always be on finding the best match for the wall, the nail and the hammer and be willing to change, improve and create.